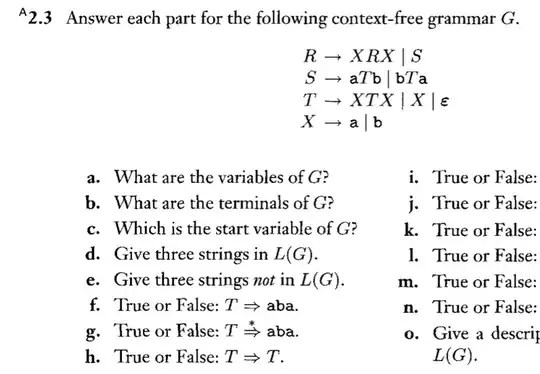
I found out that %in% stands for matching operator, binary (in model formulae: nesting). There are two tables in my workspace. The first table contains
> str(GP.drugs)
'data.frame': 4158393 obs. of 9 variables:
$ SHA : Factor w/ 10 levels "Q30","Q31","Q32",..: 1 1 1 1 1 1 1 1 1 1 ...
$ PCT : Factor w/ 151 levels "5A3","5A4","5A5",..: 16 16 16 16 16 16 16 16 16 16 ...
$ PRACTICE: Factor w/ 10191 levels "A81001","A81002",..: 344 345 345 345 345 345 345 345 345 345 ...
$ BNF.CODE: Factor w/ 1731 levels "0101010C0","0101010E0",..: 878 4 9 11 17 22 25 26 27 28 ...
$ BNF.NAME: Factor w/ 1524 levels "Abacavir ",..: 317 289 294 1284 37 379 655 825 1115 824 ...
$ ITEMS : int 1 27 1 2 97 4 40 98 27 2 ...
$ NIC : num 1.89 74.94 3.2 7.35 439.83 ...
$ ACT.COST: num 1.77 69.92 2.98 6.84 408.43 ...
$ PERIOD : num 201109 201109 201109 201109 201109 ...
The second table contains
> str(problem.drugs)
'data.frame': 13 obs. of 2 variables:
$ Drug : Factor w/ 13 levels "Alogliptin","Glipizide",..: 1 2 3 9 10 11 12 13 4 7 ...
$ Category: Factor w/ 1 level "metformin": 1 1 1 1 1 1 1 1 1 1 ...
The code and the error I am using is
> t<-subset(GP.drugs,n %in% p)
> t
[1] SHA PCT PRACTICE BNF.CODE BNF.NAME ITEMS NIC ACT.COST PERIOD
<0 rows> (or 0-length row.names)
More errors
Does it make difference on the tables' column names or does it make it difference on the number of columns both have?