Here is what it looks like after those edits - lines but no boxes.
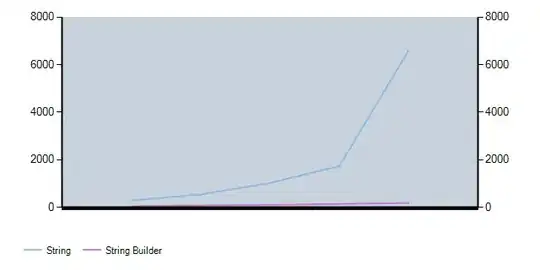
Reproducible code:
df <- data.frame(SampleID = structure(c(5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L, 5L),
.Label = c("C004", "C005", "C007", "C009", "C010",
"C011", "C013", "C027", "C028", "C029",
"C030", "C031", "C032", "C033", "C034",
"C035", "C036", "C042", "C043", "C044",
"C045", "C046", "C047", "C048", "C049",
"C058", "C086"), class = "factor"),
Sequencing.Depth = c(1L, 2612L, 5223L, 7834L, 10445L, 13056L, 15667L, 18278L,
20889L, 23500L),
Observed.OTUs = c(1, 213, 289.5, 338, 377.8, 408.9, 434.4, 453.8, 472.1, NA),
Mange = structure(c(2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L, 2L),
.Label = c("N", "Y"), class = "factor"),
SpeciesCode = structure(c(3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L, 3L),
.Label = c("Cla", "Ucin", "Vvu"), class = "factor"))